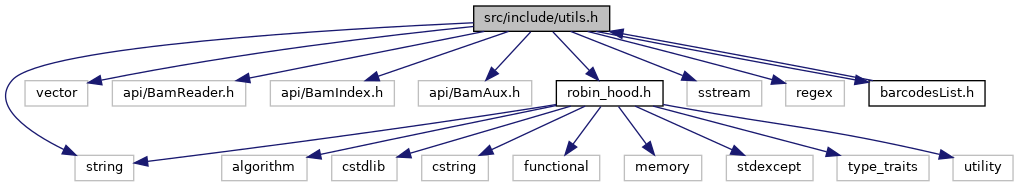
#include <string>
#include <vector>
#include "api/BamReader.h"
#include "api/BamIndex.h"
#include "api/BamAux.h"
#include "robin_hood.h"
#include <sstream>
#include <regex>
#include "barcodesList.h"
Go to the source code of this file.
◆ BXTAG
◆ no_argument
◆ optional_argument
| #define optional_argument 2 |
◆ required_argument
| #define required_argument 1 |
◆ barcode
◆ SequencingTechnology
Supported sequencing technologies
| Enumerator |
|---|
| Undefined | |
| TenX | |
| Haplotagging | |
| TELLSeq | |
| stLFR | |
Definition at line 28 of file utils.h.
◆ convertToSam()
| string convertToSam |
( |
const BamAlignment & |
a, |
|
|
RefVector |
m_references |
|
) |
| |
Translate a BamAlignment to a SAM-like string.
- Parameters
-
| a | BamAlignment to translate |
| m_references | vector containing the information (name and length) about reference sequences |
- Returns
- a SAM-like string summarizing the information of a
◆ determineSequencingTechnology()
Determine the sequencing technology the barcode originates from. This function is compatible with 10x Genomics, Haplotagging, TELL-SEq and stLFR. Barcodes that do not come from these technologies and are not represented as a suite of nucleotides will cause the program to exit.
- Parameters
-
| barcode | the barcode to determine sequencing technology from |
- Exceptions
-
| runtime_error | thrown if a barcode pattern could not be converted to a regexp or if the used sequencing technology was not recognized |
- Returns
- The SequencingTechnology enum field corresponding to the sequencing technology
◆ extractRegions()
| vector<string> extractRegions |
( |
string |
chromosome, |
|
|
int32_t |
chromosomeSize, |
|
|
unsigned |
regionSize |
|
) |
| |
Extract all regions of a given size from a civen chromosome.
- Parameters
-
| chromosme | chromosome of interest |
| chromosomeSize | size of the chromosome |
| regionSize | size of the regions to extract |
- Returns
- a list of all regions of specifed size of the chromosome
◆ extractRegionsList()
| vector<string> extractRegionsList |
( |
BamReader & |
reader, |
|
|
unsigned |
regionSize |
|
) |
| |
Extract all regions from all chromosomes.
- Parameters
-
| reader | BamReader open on the desired BAM file |
| regionSize | size of the regions to extract |
- Exceptions
-
| runtime_error | thrown if a contig name could not be converted to an ID or if a region of redaer could not be jumped to |
- Returns
- a list of all regions of specified size of all the chromosomes
◆ isValidBarcode()
| bool isValidBarcode |
( |
const string & |
barcode | ) |
|
Check whether a barcode is valid or not. A barcode is considered as valid if it is not empty, if it does not contain any "N" for 10x and TELL-Seq, if it is not "0_0_0" for stLFR data, and does not contain a "00" substring for Haplotagging data. The function takes care of determining the employed sequencing technoly.
- Parameters
-
| barcode | the barcode to verify |
- Exceptions
-
| runtime_error | thrown if the sequencing technology could not be recognized |
- Returns
- true if the barcode is valid, false otherwise
◆ retrieveNucleotidesContent()
| string retrieveNucleotidesContent |
( |
const string & |
barcode | ) |
|
Retrieve the nucleotides content of the barcode. This function is used to translate barcodes represented as a suite of integers (as in stLFT and Haplotagging) into nucleotides barcodes. The function takes care of determining the employed sequencing technoly.
- Parameters
-
| barcode | the barcode to retrieve nucleotides for |
- Exceptions
-
| runtime_error | thrown if the sequencing technology could not be recognized |
- Returns
- the barcode in nucleotides representation
◆ splitString()
| vector<string> splitString |
( |
string |
s, |
|
|
string |
delimiter |
|
) |
| |
Split a string according to a delimiter.
- Parameters
-
| s | string to split |
| delimiter | delimiter |
- Returns
- a vector containing the splits of the string
◆ stringToBamRegion()
| BamRegion stringToBamRegion |
( |
BamReader & |
reader, |
|
|
string |
s |
|
) |
| |
Translate a string to a BamRegion.
- Parameters
-
| reader | BamReader open on the desired BAM file |
| s | string to translate, formatted as chromosome:startPosition-endPosition |
- Exceptions
-
| runtime_error | thrown if a region could not be converted to a BamRegion or if a contig name could not be converted to an ID |
- Returns
- the BamRegion coressponding to the string s
◆ stringToBarcode()
| barcode stringToBarcode |
( |
const string & |
str | ) |
|
Translate a string to a barcode in 2 bits per nucleotide format. The function takes care of determining the employed sequencing technoly, and of retrieving the nucleotides contents of barcodes represented as a suite of integers.
- Parameters
-
- Returns
- the barcode in binary representation
◆ techno
 1.8.17
1.8.17